This is an archival copy of the Visualization Group's web page 1998 to 2017. For current information, please vist our group's new web page.
Computational Chemistry Grand Challenge
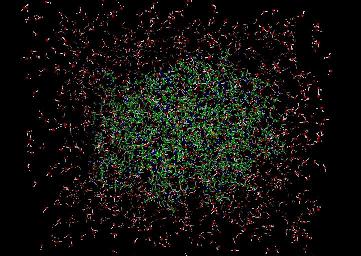 |
Researchers at Northwestern University are studying the enzyme beta-lactamase. Specifically, the research is focused upon uncovering the specific molecular mechanisms employed by the enzyme to hydrolyse penicillin G, thus rendering it biologically inactive.
This visualization shows 250 frames from a molecular dynamics simulation run on the Cray T3E at NERSC. The simulation computes the position of some 20,000 atoms at 2 femtosecond time steps, storing their coordinates to a "trajectory" file every 100 femtoseconds. The 250 frames in this visualization span 25 picoseconds of elapsed simulation time. The visualization depicts approximately 8000 atoms of the complete set of 20,000.
On the T3E, the trajectory file was converted from CHARMM format in native T3E binary format to a more general format. Next, a PDB reader was modified so as to read atomic positions from the converted CHARMM data. For the purposes of visualization, the entire converted CHARMM file was processed, one time step at a time, then rendered as a "ball and stick" model.
250 Frame MPEG movie (5.5Mbytes)
Acknowledgement
- Paul Bash, Mark Cunningham, Northwestern University