This is an archival copy of the Visualization Group's web page 1998 to 2017. For current information, please vist our group's new web page.
Drosophila Gene Expression Data Exploration and Visualization
Table of Contents
- Introduction
- What is gene expression and why is it important
- Data- and Visualization Pipeline
- Previous Work
- PointCloudXplore: Interactive 3D Visualization
- 3D/2D Embryo Views
- 3D Gene Expression Surface Graphs
- Cell Magnifier
- 3D/2D Scatterplots
- 3D Parallel Coordinates
- Brushing & Linking
- Data Clustering
- Multi View: Visualization of Embryo-Registration
- Discussion and Next Steps
- Publications
- References
Introduction
In a joint effort with the
Berkeley
Drosophila Transcription Network (BDTNP) we have developed
PointCloudXplore, a tool aimed at helping biologists to understand the
relationship between gene expression patterns in three dimensions.
The BDTNP produces novel 3D point cloud data sets, which are created
from 3D confocal microscopy images, that contain information about gene
expression in fruit fly embryos at cellular resolution. To support
analysis of these high dimensional data sets, PointCloudXplore
integrates multiple views to ease analysis of complex gene expression
data. Each view emphasizes different data properties, and interaction
between the views makes it possible to perform detailed analyses of the
presented data. This type of interaction blends high-dimensional
information exploration with interactive, 3D visualization.
Along with the BDTNP's 3D Gene Expression database we have also made PointCloudXplore available for download free of charge. PointCloudXplore is available for Linux, Windows, and MacOS. For more information see the BDTNP webpage.
What is gene expression and why is it important
The genetic information needed to create and maintain an organism is stored in strands of deoxyribonucleic acid (DNA). The DNA itself is subdivided into functional subregions, the genes. Genes themselves are not responsible for performing any function in a cell. Instead, they are coding for proteins. The transcription process copies genes into mRNA. Subsequently, the translation process uses the genetic information of the gene, which is now available in the mRNA, to produce protein. Therefore, it is possible to define gene expression as the amount of protein produced using information stored in a gene. Proteins are involved in practically every function performed by a cell, e.g., as enzymes, structural proteins or as regulatory proteins which are responsible for regulation of gene expression. In this way complex genetic regulatory networks are build up. Genetic regulatory networks are also responsible for guiding the developmental process of any organism. The goal of the BDTNP is to decipher how the patterns of gene expression underlying animal development are directed by the regulatory information contained in DNA sequences. To achieve this goal the BDTNP has chosen the fruitfly (Drosophila melanogaster) as a model organism. FlyMove provides more detailed information about the development of Drosophila melanogaster.
Data- and Visualization Pipeline
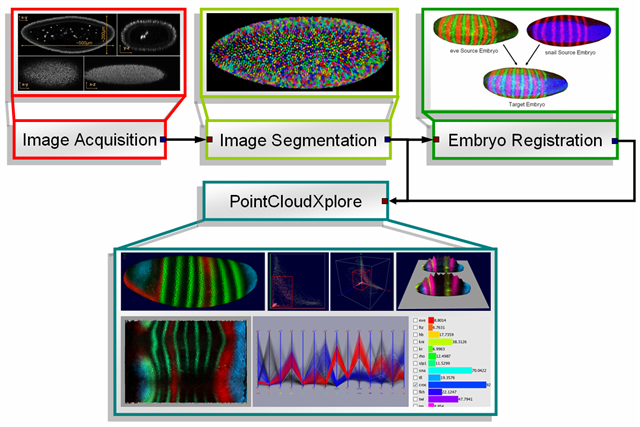
A Single PointCloud file contains information about the x, y, z location of each nucleus in an embryo, the nuclear and cytoplasmic volumes, and relative concentrations of gene products (mRNA or protein) associated with each nucleus and surrounding cytoplasm. To generate Single PointClouds, embryos are first labeled with fluorophore to detect two gene products (typically for 2 genes) and with an additional label to detect the nuclei. Using laser scanning microscopy 3D image stacks containing the whole embryo are created for each embryo. These images are then processed to detect the blastoderm nuclei and measure the expression level for the measured genes.
Due to the limited number of different spectrally distinguishable fluorophore and difficulties of adding multiple labels to an embryo it is experimentally not practical to obtain the expression more than a few genes in a single embryo. In order to understand the complex relationships between genes it is, however, critical to compare the expression regulators and their many target genes in a common coordinate framework. Therefore, a set of Single PointClouds is registered into one ore more Virtual PointClouds using morphology as well as a common reference gene to determine cell correspondences. A Virtual PointCloud contains averaged expression levels of many genes mapped onto the nuclei of a reference embryo. Using spatial as well as temporal registration the BDTNP created an expression Atlas for stage 5 of embryo development containing information of the expression of around 100 genes at up to 6 time-steps. PCX is used for visualization of both Single PointClouds and Virtual PointClouds.
| Figure 1: Data- and Visualization Pipeline |
Previous Work
Up to now studies of animal gene expression patterns have not captured 3D context. The 3D point cloud datasets described above contain information about gene expression at cellular resolution for whole fruitfly embryos. Such information has never been available before in such detail and quality. Available visualization tools are not sufficient for comparing and analyzing the generated 3D PointCloud datasets. PointCloudXplore is a tool specially designed for visualization of 3D gene expression data in early stage embryos of Drosophila melanogaster. Many different views and interactive interaction with the data open a way to interactive data analysis like it was never possible before in this specific area of research. To improve and facilitate the data analysis process we have integrated data into PointCloudXplore. The interplay of data clustering and visualization improves the visualization as well as the clustering process.
PointCloudXplore: Interactive 3D Visualization
PointCloudXplore is based on two simple but powerful basic principles. Multiple views are used to view the data from different perspectives without being overwhelmed by the high dimensionality of the data. Each view emphasizes different data properties and the interplay between all views makes detailed data analysis possible. The second basic principle is called Brushing&Linking. Brushing refers to the fact that the user can select cells of interest according to different data properties in any view. Cells selected in one view are highlighted visually in all other data displays. In this way all views are linked together. Linking simply means that it is possible to identify visually which parts of one data display correspond to that of another one.
3D/2D Embryo Views
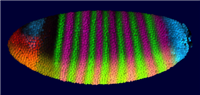
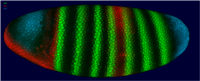
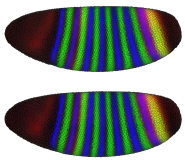
In PointCloudXplore we use several models of the embryo to enable analysis of spatial gene expression pattern. Each cell is represented by one 3D graphical object (sphere or polygon) positioned in space according to the physical position of the cell it represents. We use a simulated color staining of the embryo model to visualize gene expression values. For each gene the user selects a basic color while gene expression level are mapped to color brightness. Besides 3D models of the embryo we also make use of 2D embryo representation to make it possible to look at all cells in parallel. Simply by drawing on the embryo surface the user can select cells of interest with respect to their position in physical space. Cells selected by the user highlighted again using color.
| Figure2: Sphere View | Figure3: Cell View | Figure4: Unrolled View | Figure 5: 2D Projection View |
3D Gene Expression Surface Graphs
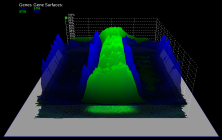
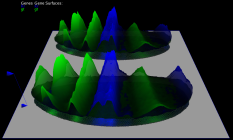
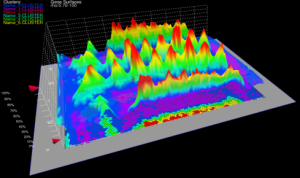
3D graphs, defined over the 2D embryo views, allow qualitative and quantitative analysis of gene expression data. The cell position in the underlying 2D embryo view determines the x/y-position of surface points, whereas height of a gene expression surface corresponds to the expression values measured for the gene it represents. Looking at several gene expression offset surfaces at once reveals relationships between genes.
| Figure 6: 3D Offset surfaces on Unrolled View | Figure 7: 3D Offset surfaces of gt and Kr on orthographic projection of the embryo |
Cell Magnifier
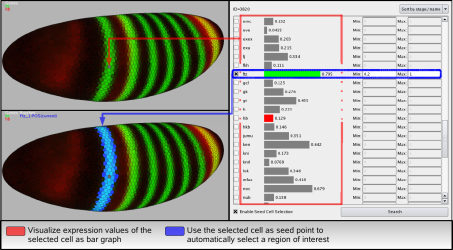
By selecting one cell in an embryo view it is possible to view all expression values measured in this cell as bar graph in the Cell Magnifier. By comparing graphs of different cells it is possible to identify some general behavior. Using this information it is then directly possible to select cells automatically according to user-defined ranges in gene expression. Using the cell displayed in the Cell Magnifier as seed point for the selection process one can fast and easily select cells in a contiguous region on the embryo that suffices the user defined expression criteria.
| Figure 8: Cell Magnifier and seed cell selection |
3D/2D Scatter-plots
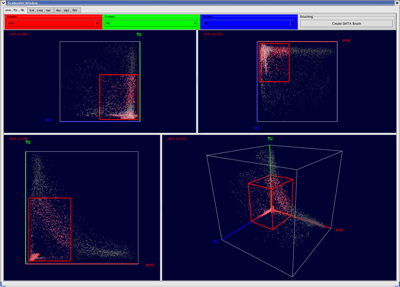
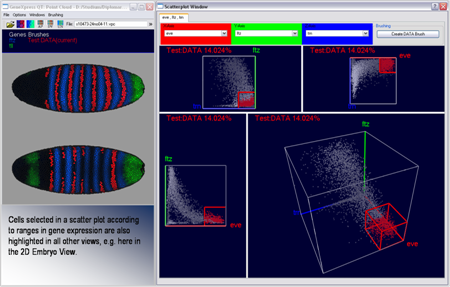
Conceptually scatter-plots represent the simplest–but at the same time a very effective way–to visualize gene inter-relationships directly in gene expression space. In a Cartesian coordinate system, which serves as reference frame, each axis represents the expression of one selected gene ranging from 0% expression at the origin to 100% relative expression. A single data point, positioned in the plot according to the relative expression levels measured, represents each cell. In PointCloudXplore we make use of halos, depth coloring, and alpha blending to improve the visualization of scatter-plots. By looking at the distribution of points in a scatter-plot, characteristic relationships between genes can be seen. Cells of interest can be selected in scatter-plots simply by drawing a box in the plot. It is possible to analyze the spatial pattern of cells selected in a scatter-plot in any physical view. The interplay of visualizations in physical and gene expression space enables in this way detailed analysis of the data.
| Figure 9: 2D and 3D Scatterplots | Figure 10: Scatterplots and Embryo View in interaction |
3D Parallel Coordinates
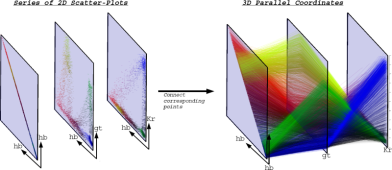
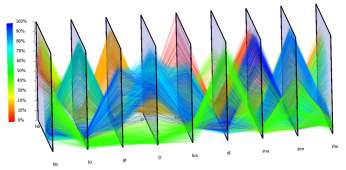
To enable visualization of the expression of many genes in parallel, we adapted parallel coordinates to visualization of 3D gene expression data. In parallel coordinates, each gene is represented by one parallel axis. The expression levels of a cell define a point on each axis. By connecting the corresponding points of neighboring axes, each cell can be represented by a polyline.
Spatial information is crucial in the analysis of 3D gene expression. Information about the spatial relationships between different genes' expression patterns is essential for the analysis of regulatory networks. To display this information in parallel coordinate Views, we extrude the coordinate axes into the third dimension and order the data lines–each representing one cell–back-to-front according to their position along the AP- or DV axis of the embryo. Using this 3D visualization, spatial and gene expression information are clearly separated while the basic character of spatial gene expression patterns is preserved in one dimension. Besides spatial information we also support display of gene expression information along the third dimension. Cell of interest can be selected by defining ranges in gene expression using two sliders attached to each axis.
 |
 |
|
| Figure 11: From 2D scatter-plots to 3D parallel coordinates | Figure 12: 3D parallel coordinate view of all cells and nine selected genes |
Brushing & Linking
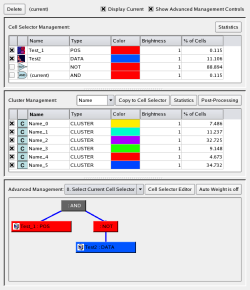
In PointCloudXplore, we store all cell selections in a central Cell Selector Management system to effectively link all view together. Via this central management, all views have access to the same set of cell selections so that cells selected in any view are also highlighted in any other view. Using logical operation, such as AND, OR, and NOT, the user can define advanced cell queries by combining individual cell selection from any view. These logical operators are again implemented as cell selection allowing one to define complex logical selection models that can be represented as logical trees.
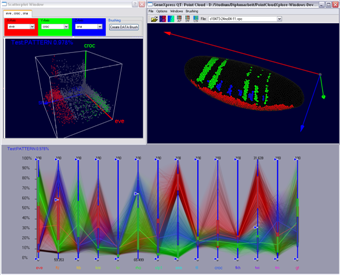
| Figure 13: Management of Cell Selectors and Clusters | Figure 14: Parallel coordinates in interaction with a 3D scatter-plot and the 3D embryo view |
Data Clustering
To facilitate the analysis process, we use data clustering to identify groups of cells that behave similar with respect to the expression of selected genes. Data clustering can, e.g., be used to classify the expression pattern of a single gene by creating a set of data-dependent thresholds. Groups of cells with similar temporal expression behavior can be identified by considering the expression of the same gene at several time-points in the cluster analysis. Data clustering can, in this way, be used to effectively analyze the temporal variation of a gene expression pattern. Besides for the analysis of expression characteristics of single genes, we have also used data clustering for validation of gene inter-relationships. Characteristic groups of cells behaving similar with respect to the expression of suspected regulators can be identified by considering the expression of several different genes of interest in the clustering process. By comparing the patterns formed by these clusters with the pattern of potential target genes hypothesis of how a pattern of a gene is defined by its regulators can be defined.
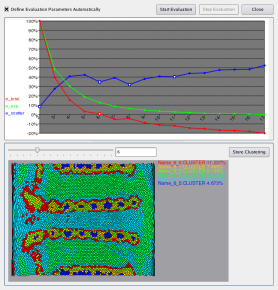
In may cases an appropriate number of clusters may be unknown. To assist the user in identifying a well suited number of clusters, we use specifically developed cluster quality measures. As shown in Figure 14, by iterating over the number of clusters and calculating: i) the physical scattering (blue); ii) the error in expression space (green); and iii) the total error (red); the users can effectively evaluate an appropriate number of clusters.
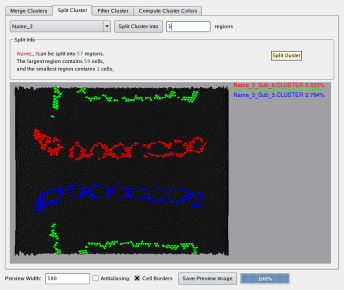
In order to allow effective validation of clustering results dedicated statistics plots are made available in PointCloudXplore. Using dedicated post-processing techniques the user can create finer or coarser representations of the initial clustering results as well as improve the clustering results. Manual correction and filtering of clusters based on spatial information are used for improvement of clustering results. Merging of clusters and splitting of clusters into their main spatial components (see Figure 15) are used to derive coarser and finer representations from an initial clustering.
Like a user defined Cell Selector, an automatically created Cluster simply defines a selection of cells. Clusters can therefore be managed (see Figure 12) and visualized in the same way as user defined cell selections. In this way, parallel coordinates, scatter-plots as well physical views of the embryo can be used for analysis of clustering results. Figure 16, e.g., shows how data clustering is used for classification of the rho expression pattern. Clustering results are then mapped on an expression surface of rho.
Discussion and Next Steps
Future work will concentrate on development of tools for comparative visualization of different time-steps of embryo development as well as of wild-type and mutant data. Analysis of the temporal variation of gene expression pattern is essential in order gain insight in the complex dynamic behavior of gene expression. By comparing data from wild-type and mutant embryos it is possible to gain a deeper understanding of the function of different genes and their interactions with other genes. Integration of further tools for data analysis will be another main focus of our future work. Tools that enable one to derive hypotheses of potential interactions between genes –that can then be tested using, e.g., in-vivo and in-vitro binding data–are essential for the efficient and effective analysis of the complex genetic regulatory networks that guide embryo development.
[1] O. Rübel, G. H. Weber, M-Y Huang, E. W. Bethel, M. D. Biggin, C. C. Fowlkes, C. Luengo Hendriks, S. V. E. Keränen, M. Eisen, D. Knowles, J. Malik, H. Hagen and B. Hamann, "Integrating Data Clustering and Visualization for the Analysis of 3D Gene Expression Data." IEEE Transactions on Computational Biology and Bioinformatics. 2008. LBNL-382E. (PDF), (BibTeX)
[2] G. H. Weber, O. Rübel, M.-Y. Huang, A. H. DePace, C. C. Fowlkes, S. V. E. Keränen, C. L. Luengo Hendriks, H. Hagen, D. W. Knowles, J. Malik, M. D. Biggin and B. Hamann. "Visual exploration of three-dimensional gene expression using physical views and linked abstract views." In IEEE Transactions on Computational Biology and Bioinformatics. LBNL-63776. (PDF), (BibTeX).
[3] M.-Y. Huang, O. Rübel, G.H. Weber, C.L. Luengo Hendriks, M.D. Biggin, H. Hagen, B. Hamann. "Segmenting Gene Expression Patterns of Early-stage Drosophila Embryos.", In: L. Linsen, H. Hagen, B. Hamann, eds., Mathematical Methods for Visualization in Medicine and Life Sciences. Springer-Verlag, Heidelberg, Germany, 2008. LBNL-62450. (PDF), (BibTeX).
[4] O. Rübel, G.H. Weber, S.V.E. Keränen, C.C. Fowlkes, C.L. Luengo Hendriks, L. Simirenko, N.Y. Shah, M.B. Eisen, M.D. Biggin, H. Hagen, J.D. Sudar, J. Malik, D.W. Knowles, and B. Hamann. PointCloudXplore: Visual analysis of 3D gene expression data using physical views and parallel coordinates. In: B. Sousa Santos, T. Ertl, and K.I. Joy, eds., Data Visualization 2006 (Proceedings of EuroVis 2006), Eurographics Association, Aire-la-Ville, Switzerland, pp. 203-210. LBNL-60005.(BibTeX)
References
[1] C.L. Luengo Hendriks*, S.V.E. Keraenen*, C.C. Fowlkes, L. Simirenko, G.H. Weber, A.H. DePace, C.N. Henriquez, D.W. Kaszuba, B. Hamann, M.B. Eisen, J. Malik, D. Sudar, M.D. Biggin and D.W. Knowles, 3D morphology and gene expression in the Drosophila blastoderm at cellular resolution I: data acquisition pipeline, Genome Biology, 7:R123, December 2006. (available online at http://genomebiology.com/2006/7/12/R123 )
[2] S.V.E. Keraenen*, C.C. Fowlkes*, C.L. Luengo Hendriks*, D. Sudar, D.W. Knowles, J. Malik and M.D. Biggin, 3D morphology and gene expression in the Drosophila blastoderm at cellular resolution II: dynamics, Genome Biology, 7:R124, December 2006. ( available online at http://genomebiology.com/2006/7/12/R124 )
[3] Weber G. H. , Ruebel O., Huang M.-Y., DePace A. H., Fowlkes C. C., Ker�nen S. V. E., Luengo Hendriks C. L., Hagen H., Knowles D. W., Malik J., Biggin M. D., and Hamann B. , Visual exploration of threedimensional gene expression using physical views and linked abstract views, Accepted for Publication in IEEE Transactions on Computational Biology and Bioinformatics, 2007.
[4] Fowlkes, C.C., Luengo Hendriks, C.L., Keraenen, S.V.E., Weber, G.H., Ruebel, O., Huang, M.-Y., Chatoor, S., Simirenko, L., Henriquez, C., Beaton, A., Weiszmann, R., Celniker, S., Eisen, M.B., Hamann, B., Knowles, D.W., Biggin, M.D. and Malik, J., Constructing a quantitative spatio-temporal atlas of gene expression in the Drosophila blastoderm, in review.
[5] G.H. Weber, C.L. Luengo Hendriks, S.V.E. Keraenen, E.Dillard, D.Y. Ju, D.Sudar, and B. Hamann. Visualization for Validation and Imporvement of Three-dimensional Segmentation Algorithms. EUROGRAPHICS- IEEE VGTC Symposium on Visualization 2005.
[6] Harald Piringer, Robet Korsara, Helwig Hauser. Interactive Focus+Context Visualization with Linked 2D/3D Scatterplots. 2nd International Conference on Coordinated and Multiple Views in Exploratory Visualization (CMV), 2004.
[7] Robert Krosara, Gerald N. Sahling, Helwig Hauser. Linking Scientific and Information Visualization with Interactive 3D Scatterplots. Short Communication Papers Proceedings of the 12th International Conference in Central Europe on Computer Graphics, Visualization, and Computer Vision (WSCG), pp. 133-140, 2004.
[8] Alfred Inselberg. Visualizing High Dimensional Datasets \& Multivariate Relations. Tutorials for KDD 2000. University of Tel Aviv, Israel. August 20, 2000.
[9] A. Inselberg and B. Dimsdale. Parallel coordinates: A tool for visualizing multidimensional geometry. Proceedings of the 1st IEEE Conference on Visualization (Vis '90), pages 361--378, 1990.
[10] Ying-Huey Fua , Matthew O. Ward , Elke A. Rundensteiner. Hierarchical parallel coordinates for exploration of large datasets. IEEE Visualization, Proceedings of the conference on Visualization '99. San Francisco, California, United States Pages: 43 - 50. 1999 ISBN:0-7803-5897.
[11] Helwig Hauser, Florian Ledermann, and Helmut Doleisch. Angular Brushing of Extended Parallel Coordinates. INFOVIS, Proceedings of the IEEE Symposium on Information Visualization (InfoVis'02), Page: 127. 2002 ISBN:0-7695-1751-X.
[12]. Visually Effective Information Visualization of Large Data. Matej Novotny VRVis Research Center for Virtual Reality and Visualization. Vienna / Austria. 2004.
[13] Jing Yang, Wei Peng, Matthew O. Ward and Elke A. Rundensteiner. Interactive Hierarchical Dimension Ordering, Spacing and Filtering for Exploration of High Dimensional Datasets. IEEE Symposium on Information Visualization 2003 (InfoVis 2003), pp 105 - 112, October 2003.
[14] Jing Yang, Matthew O. Ward, Elke A. Rundensteiner and Shiping Huang, "Visual Hierarchical Dimension Reduction for Exploration of High Dimensional Datasets", VisSym 2003.
[15] M.B.
Eisen, P.T. Spellman, P.O. Brown and D.
Botstein. Cluster Analysis and Display of Genome-Wide
Expression Patterns. Proc Natl Acad Sci U S.
Page: 14863-14868. 1995
[16] J. Handl, J. Knowles and D. B. Kell. Computational cluster validation in post-genomic data analysis. Bioinformatics. Vol 21. Number 152005, Pages 3201-3212. May.2005
[17] M. J. L. de Hoon, S. Imoto, J. Nolan and S. Miyano. Open Source Clustering Software. Bioinformatics. Vol 20(9). Pages 1453 - 1454. 2004
[18] A. K. Jain, M. N. Murty and P. J. Flynn. Data Clustering: A Review. ACM Computing Surveys. Vol 31(3) , Sept.1999